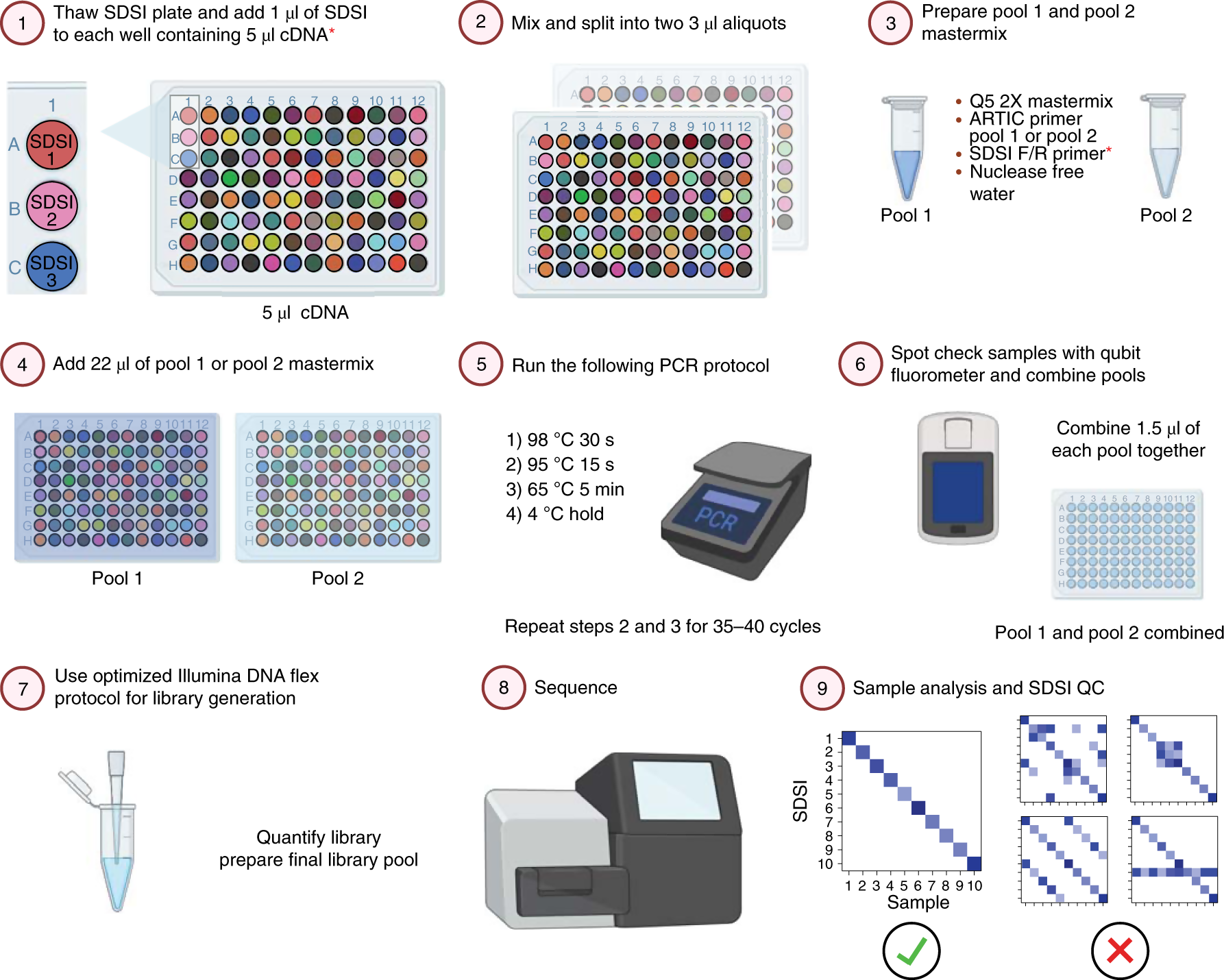
Synthetic DNA spike-ins (SDSIs) enable sample tracking and detection of inter-sample contamination in SARS-CoV-2 sequencing workflows | Nature Microbiology
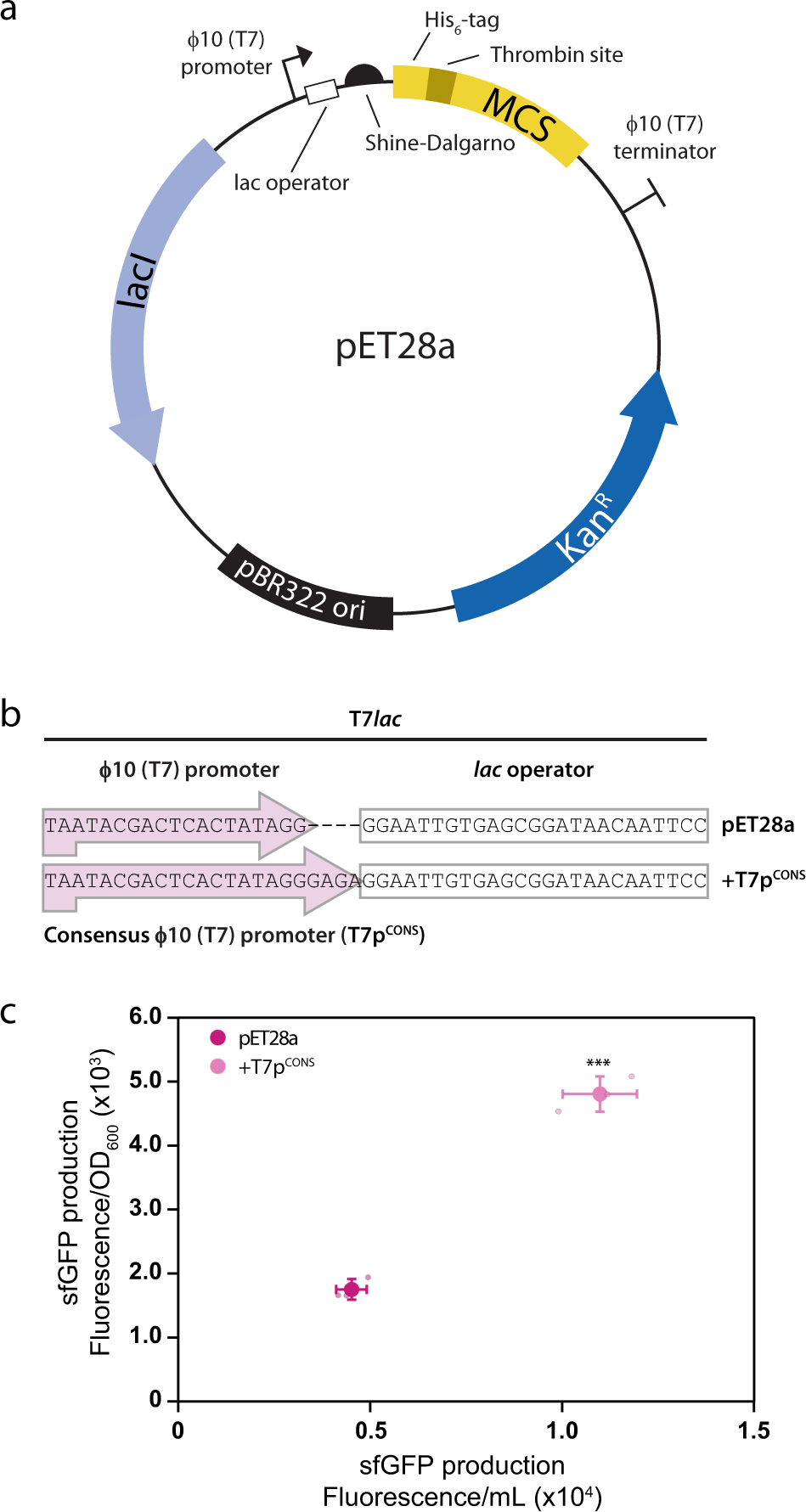
Improved designs for pET expression plasmids increase protein production yield in Escherichia coli | Communications Biology

Comprehensive Profiling of Four Base Overhang Ligation Fidelity by T4 DNA Ligase and Application to DNA Assembly | ACS Synthetic Biology

Overview of a multiplex oligonucleotide ligation-PCR (MOL-PCR) assay.... | Download Scientific Diagram

Prediction and verification of glycosyltransferase activity by bioinformatics analysis and protein engineering - ScienceDirect

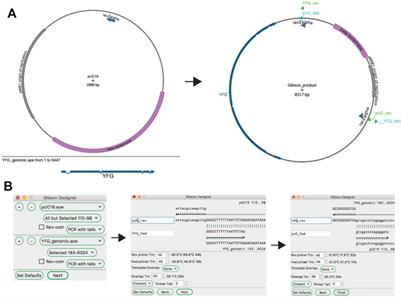
Direct Pathway Cloning Combined with Sequence- and Ligation-Independent Cloning for Fast Biosynthetic Gene Cluster Refactoring and Heterologous Expression | ACS Synthetic Biology
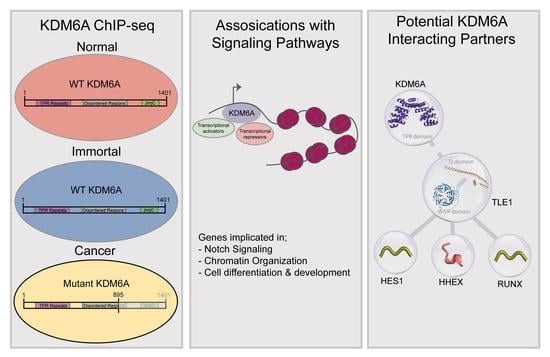
Cells | Free Full-Text | Differential Occupancy and Regulatory Interactions of KDM6A in Bladder Cell Lines

Combined nanopore and single-molecule real-time sequencing survey of human betaherpesvirus 5 transcriptome | Scientific Reports

SEVA 3.1: enabling interoperability of DNA assembly among the SEVA, BioBricks and Type IIS restriction enzyme standards - Damalas - 2020 - Microbial Biotechnology - Wiley Online Library
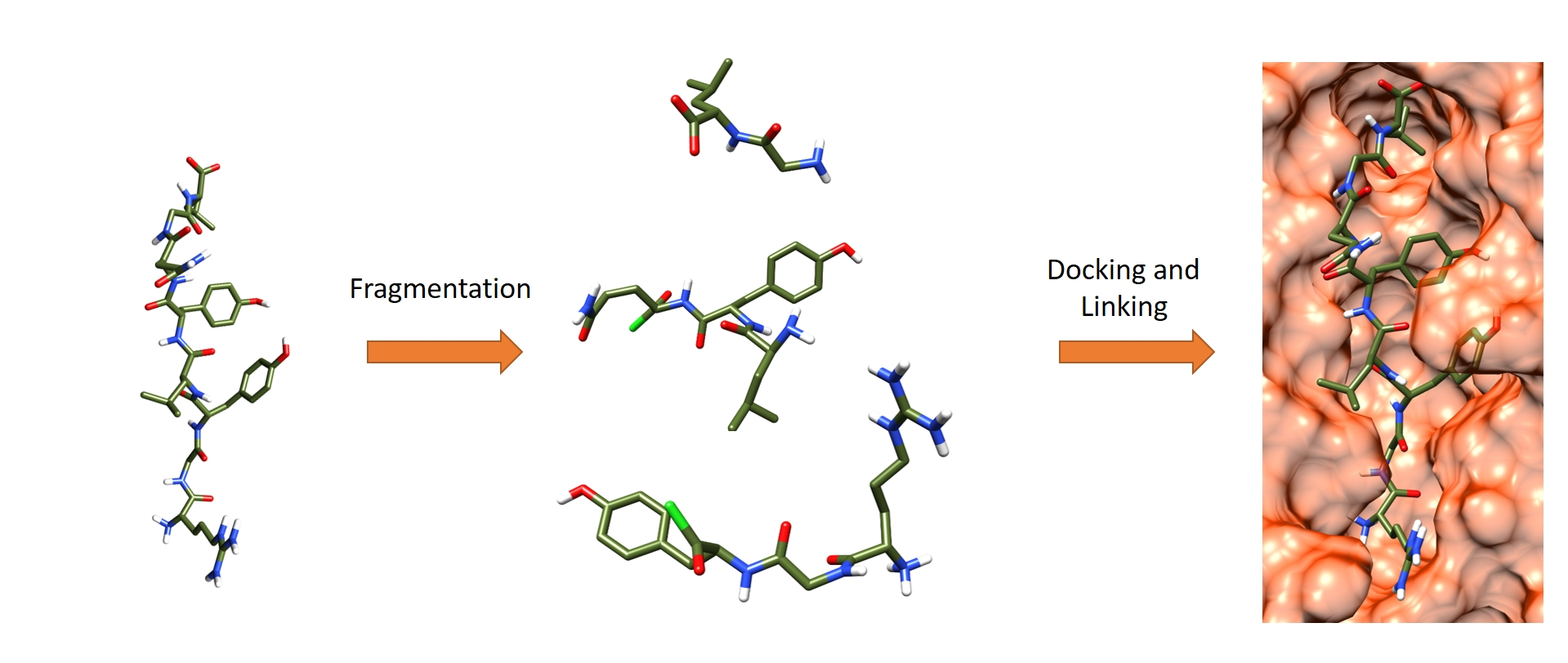
Molecules | Free Full-Text | In Silico Peptide Ligation: Iterative Residue Docking and Linking as a New Approach to Predict Protein-Peptide Interactions

Comprehensive Profiling of Four Base Overhang Ligation Fidelity by T4 DNA Ligase and Application to DNA Assembly | ACS Synthetic Biology